Kasia Kedzierska
DPhil Candidate in Genomic Medicine and Statistics
Wellcome Trust Centre for Human Genetics
University of Oxford
Hello! 👋
My research interest lies at the intersection of Machine Learning and Biology and Healthcare. The rapid advancements in technology, especially those enabling investigating genomics at single cell resolution, coupled with the progress in Machine Learning, have rendered computational biology an exceptionally captivating and invigorating field of study. Within this space, I’m particularly interested in Multimodal and Generative Models.
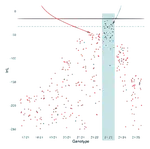
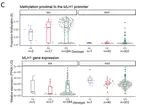
For my PhD at the University of Oxford I studied the cancer of the uterus and the role chromatin organisation plays in its initiation and progression. During my studies I interned at Novo Nordisk Research Centre in Oxford where I worked with NLP and knowledge graphs to identify potential new drug candidates.
Recently, I joined Microsoft Research New England as an intern where I am mentored by Alex Lu, Ava Amini, and Lorin Crawford. During the course of the internship I am working with Foundation Models in Computational Biology.
If you find anything on this website interesting or just fancy a chat about the intersection of biology, healthcare and Machine Learning be sure to reach out!
Actually, my full name is Katarzyna Kedzierska, or Katarzyna Zofia Kedzierska (including my middle name). In Polish, we usually have a shorter version of a name that we go by and reserve the full legal name (here: Katarzyna) for official occasions, or when we get in trouble with our parents. Here you can find recordings on how to pronounce my shorter name: Kasia or longer: Katarzyna.
- Machine Learning, Generative Models
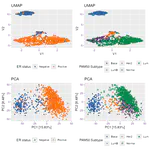
- Single cell genomics
- Cancer, Tumor heterogeneity
PhD in Genomic Medicine and Statistics, in progress
University of Oxford
MSc in Biotechnology, 2018
Warsaw University of Technology
Projects
You can read about my recent projects below. Those include my past work and DPhil project, as well as my volunteer work and hobbies.
Recent Publications
Talks and tutorials
Experience
- PhD project: Functional and evolutionary characterisation of chromatin organisation in endometrial cancer
- Together with Andrew Kwok, we are running Old Road Dhil Journal Club - computational biology and genomics JC.
- Conducted research towards Master’s thesis titled: “Analysis of the mutational burden across gene sets in cancer” defended in September 2018 at the Warsaw University of Technology
- Developed SONiCS - a tool for genotyping short tandem repeats (STRs) profiled using capture assays, SONiCS on GitHub
- Before I joined Aakrosh Ratan’s group, I worked on epigenetic regulation in prostate cancer in Pemberton group, also at UVa. By joining Ratan’s group I fully transitioned from experimental to computational biology.
Contact
- Roosevelt Drive, Oxford, Oxfordshire, OX3 7BN
- DM Me